python中将DNA链转换为RNA链
001、利用循环结构
[root@PC1 test01]# ls a.fa test.py [root@PC1 test01]# cat a.fa ## 测试DNA序列 GATGGAACTTGACTACGTAAATT [root@PC1 test01]# cat test.py ## 转换程序 #!/usr/bin/env python # -*- coding: utf-8 -*- in_file = open("a.fa", "r") str1 = "" for i in in_file: i = i.strip() for j in i: if j == "T": str1 += "U" else: str1 += j in_file.close() print(str1) [root@PC1 test01]# python test.py ## 转换结果 GAUGGAACUUGACUACGUAAAUU
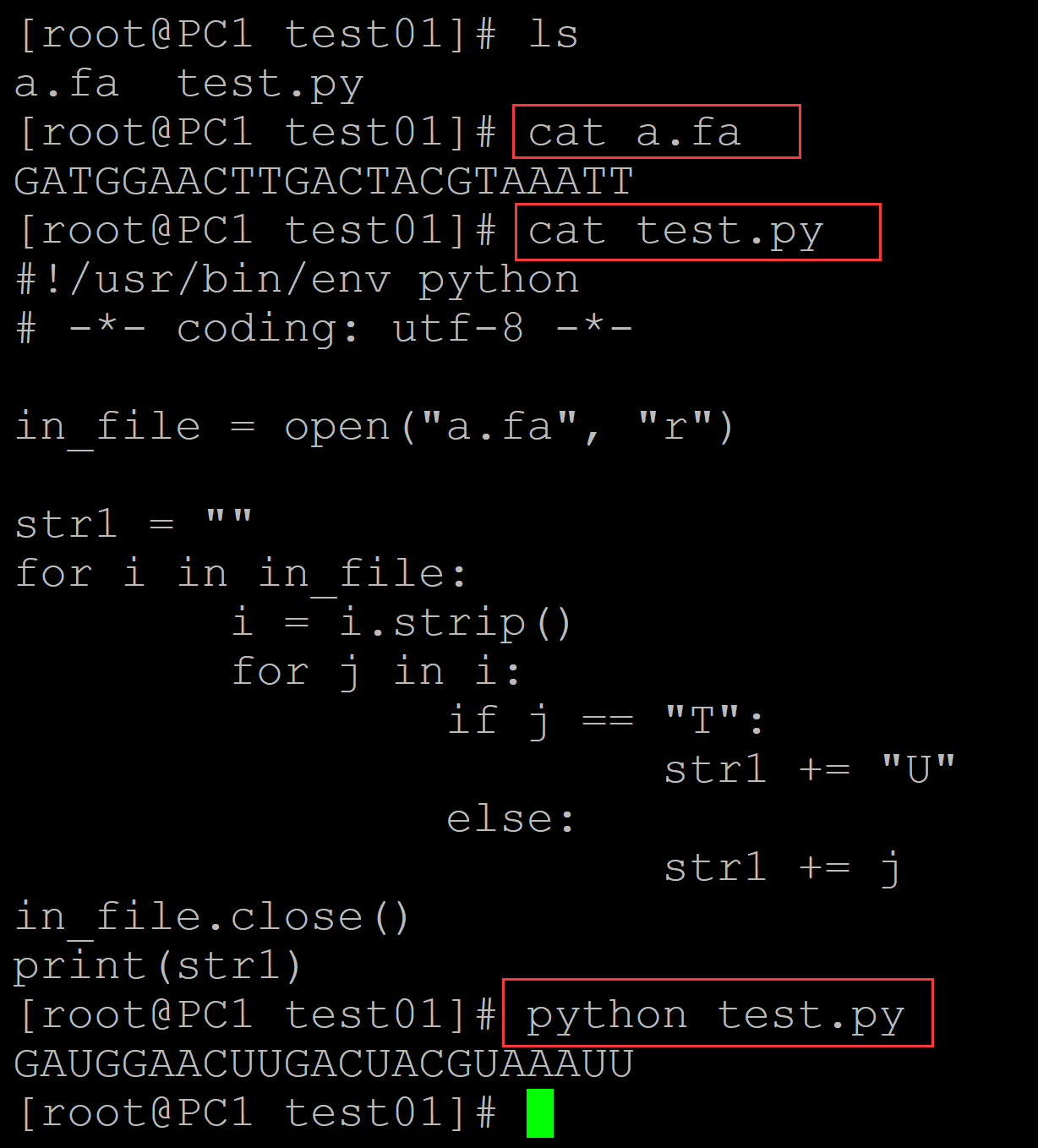
002、利用循环结构
[root@PC1 test01]# ls a.fa test.py [root@PC1 test01]# cat a.fa ## 测试DNA序列 GATGGAACTTGACTACGTAAATT [root@PC1 test01]# cat test.py ## 转换程序 #!/usr/bin/env python # -*- coding: utf-8 -*- in_file = open("a.fa", "r") file = in_file.read().strip() for i in file: if i == "T": print("U", end = "") else: print(i, end = "") in_file.close() print("") [root@PC1 test01]# python3 test.py ## 转换结果 GAUGGAACUUGACUACGUAAAUU
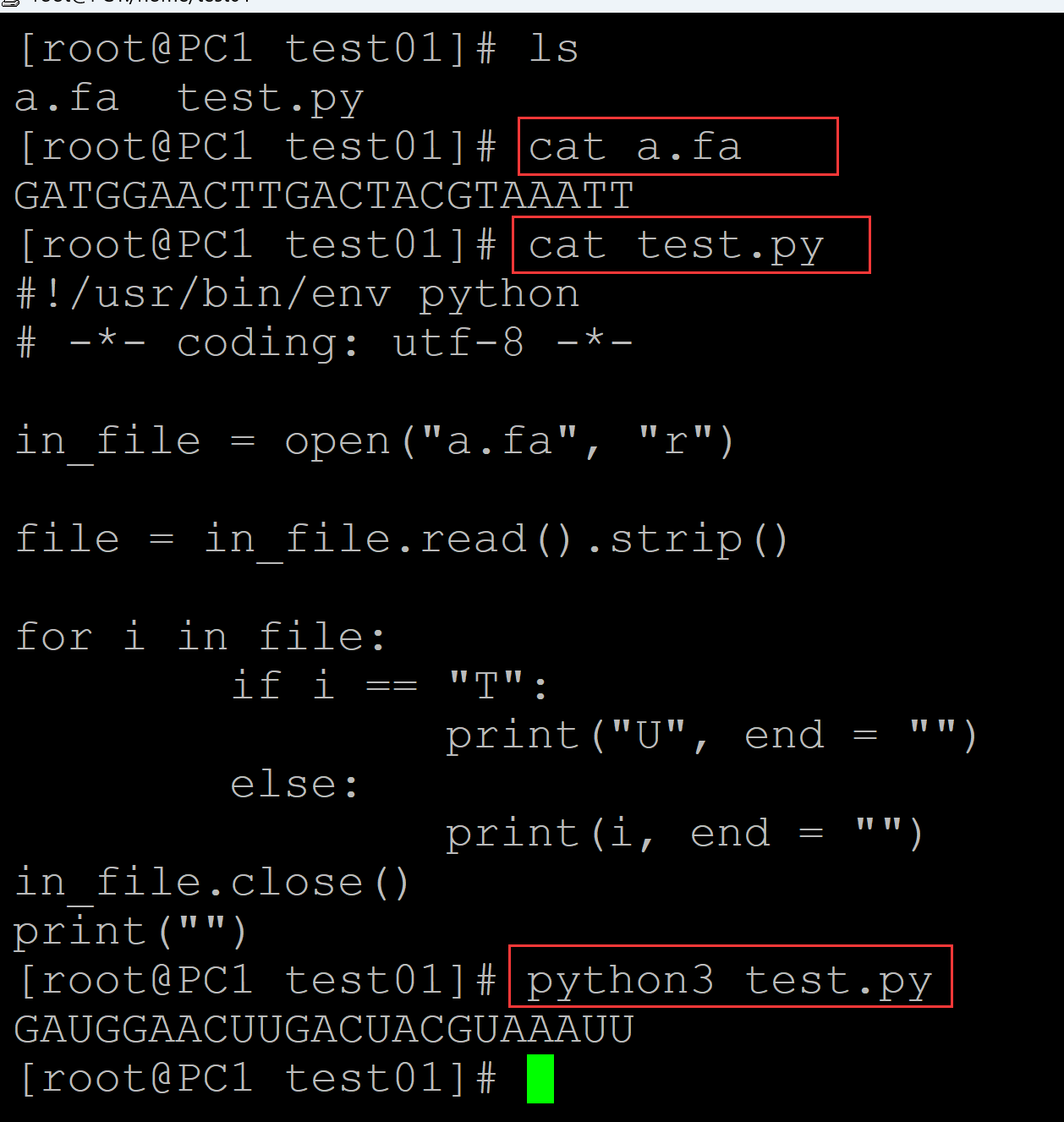
003、利用字符串替换实现
[root@PC1 test01]# ls a.fa test.py [root@PC1 test01]# cat a.fa ## 测试DNA序列 GATGGAACTTGACTACGTAAATT [root@PC1 test01]# cat test.py ## 转换程序 #!/usr/bin/env python # -*- coding: utf-8 -*- in_file = open("a.fa", "r") file = in_file.read().strip() print(file.replace("T", "U")) [root@PC1 test01]# python test.py ## 转换结果 GAUGGAACUUGACUACGUAAAUU
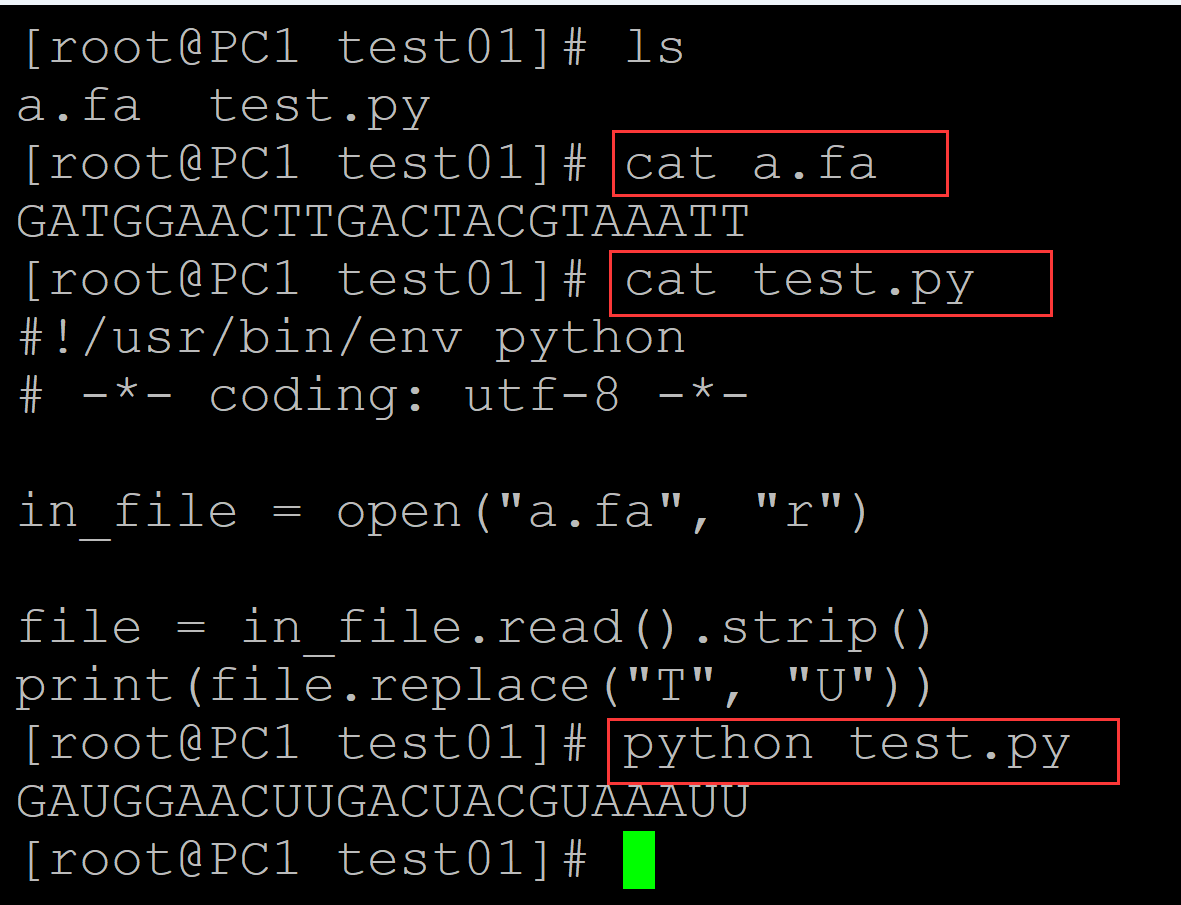
004、利用函数结构实现
[root@PC1 test01]# ls a.fa test.py [root@PC1 test01]# cat a.fa ## 测试DNA序列 GATGGAACTTGACTACGTAAATT [root@PC1 test01]# cat test.py ## 转换程序 #!/usr/bin/env python # -*- coding: utf-8 -*- in_file = open("a.fa", "r") file = in_file.read().strip() def trans(dna): return dna.upper().replace("T", "U") print(trans(dna = file)) [root@PC1 test01]# python3 test.py ## 转换结果 GAUGGAACUUGACUACGUAAAUU

参考:
https://mp.weixin.qq.com/s?__biz=MzIxMjQxMDYxNA==&mid=2247484157&idx=2&sn=869266deba638d49aa5e0bc6d2b0428c&chksm=9747cb64a03042720674040dbf19031021d298e3df300810c51883a07812419427021fd781aa&cur_album_id=1635727573621997580&scene=190#rd





【推荐】国内首个AI IDE,深度理解中文开发场景,立即下载体验Trae
【推荐】编程新体验,更懂你的AI,立即体验豆包MarsCode编程助手
【推荐】抖音旗下AI助手豆包,你的智能百科全书,全免费不限次数
【推荐】轻量又高性能的 SSH 工具 IShell:AI 加持,快人一步
· 震惊!C++程序真的从main开始吗?99%的程序员都答错了
· 【硬核科普】Trae如何「偷看」你的代码?零基础破解AI编程运行原理
· 单元测试从入门到精通
· 上周热点回顾(3.3-3.9)
· winform 绘制太阳,地球,月球 运作规律
2022-08-26 R语言中apply函数的用法
2022-08-26 R语言中colSums和rowSums函数
2022-08-26 R语言中绘制小提琴图
2021-08-26 c语言中限制用户输入整数
2021-08-26 c语言 输入验证(限制输入正数)