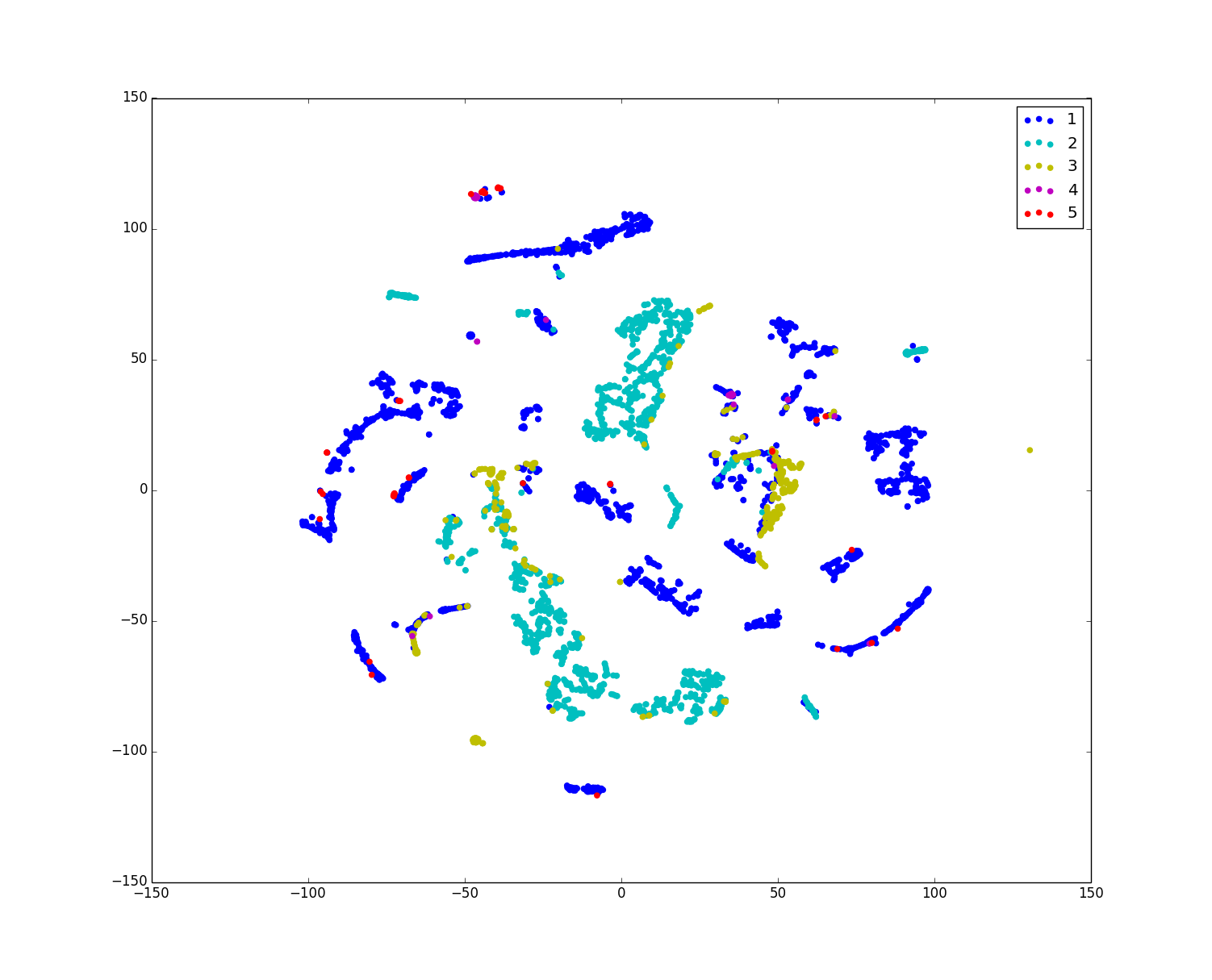
Python代码:准备训练样本的数据和标签:train_X4000.txt、train_y4000.txt 放于tsne.py当前目录.(具体t-SNE – Laurens van der Maaten http://lvdmaaten.github.io/tsne/,Python implementation),
tsne.py代码:(为了使得figure显示数据的标签,代码做了简单修改)
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 | #!/usr/bin/env python# -*- coding: utf-8 -*-## tsne.py# # Implementation of t-SNE in Python. The implementation was tested on Python 2.5.1, and it requires a working# installation of NumPy. The implementation comes with an example on the MNIST dataset. In order to plot the# results of this example, a working installation of matplotlib is required.# The example can be run by executing: ipython tsne.py -pylab### Created by Laurens van der Maaten on 20-12-08.# Copyright (c) 2008 Tilburg University. All rights reserved.import numpy as Mathimport pylab as Plot def Hbeta(D = Math.array([]), beta = 1.0): """Compute the perplexity and the P-row for a specific value of the precision of a Gaussian distribution.""" # Compute P-row and corresponding perplexity P = Math.exp(-D.copy() * beta); sumP = sum(P)+1e-6; H = Math.log(sumP) + beta * Math.sum(D * P) / sumP; P = P / sumP; return H, P; def x2p(X = Math.array([]), tol = 1e-5, perplexity = 30.0): """Performs a binary search to get P-values in such a way that each conditional Gaussian has the same perplexity.""" # Initialize some variables print "Computing pairwise distances..." (n, d) = X.shape; sum_X = Math.sum(Math.square(X), 1); D = Math.add(Math.add(-2 * Math.dot(X, X.T), sum_X).T, sum_X); P = Math.zeros((n, n)); beta = Math.ones((n, 1)); logU = Math.log(perplexity); # Loop over all datapoints for i in range(n): # Print progress if i % 500 == 0: print "Computing P-values for point ", i, " of ", n, "..." # Compute the Gaussian kernel and entropy for the current precision betamin = -Math.inf; betamax = Math.inf; Di = D[i, Math.concatenate((Math.r_[0:i], Math.r_[i+1:n]))]; (H, thisP) = Hbeta(Di, beta[i]); # Evaluate whether the perplexity is within tolerance Hdiff = H - logU; tries = 0; while Math.abs(Hdiff) > tol and tries < 50: # If not, increase or decrease precision if Hdiff > 0: betamin = beta[i].copy(); if betamax == Math.inf or betamax == -Math.inf: beta[i] = beta[i] * 2; else: beta[i] = (beta[i] + betamax) / 2; else: betamax = beta[i].copy(); if betamin == Math.inf or betamin == -Math.inf: beta[i] = beta[i] / 2; else: beta[i] = (beta[i] + betamin) / 2; # Recompute the values (H, thisP) = Hbeta(Di, beta[i]); Hdiff = H - logU; tries = tries + 1; # Set the final row of P P[i, Math.concatenate((Math.r_[0:i], Math.r_[i+1:n]))] = thisP; # Return final P-matrix print "Mean value of sigma: ", Math.mean(Math.sqrt(1 / beta)) return P; def pca(X = Math.array([]), no_dims = 50): """Runs PCA on the NxD array X in order to reduce its dimensionality to no_dims dimensions.""" print "Preprocessing the data using PCA..." (n, d) = X.shape; X = X - Math.tile(Math.mean(X, 0), (n, 1)); (l, M) = Math.linalg.eig(Math.dot(X.T, X)); Y = Math.dot(X, M[:,0:no_dims]); return Y;def tsne(X = Math.array([]), no_dims = 2, initial_dims = 50, perplexity = 30.0): """Runs t-SNE on the dataset in the NxD array X to reduce its dimensionality to no_dims dimensions. The syntaxis of the function is Y = tsne.tsne(X, no_dims, perplexity), where X is an NxD NumPy array.""" # Check inputs if X.dtype != "float64": print "Error: array X should have type float64."; return -1; #if no_dims.__class__ != "": # doesn't work yet! # print "Error: number of dimensions should be an integer."; # return -1; # Initialize variables X = pca(X, initial_dims).real; (n, d) = X.shape; max_iter = 1000 initial_momentum = 0.5; final_momentum = 0.8; eta = 500; min_gain = 0.01; Y = Math.random.randn(n, no_dims); dY = Math.zeros((n, no_dims)); iY = Math.zeros((n, no_dims)); gains = Math.ones((n, no_dims)); # Compute P-values P = x2p(X, 1e-5, perplexity); P = P + Math.transpose(P); P = P / (Math.sum(P)); P = P * 4; # early exaggeration P = Math.maximum(P, 1e-12); # Run iterations for iter in range(max_iter): # Compute pairwise affinities sum_Y = Math.sum(Math.square(Y), 1); num = 1 / (1 + Math.add(Math.add(-2 * Math.dot(Y, Y.T), sum_Y).T, sum_Y)); num[range(n), range(n)] = 0; Q = num / Math.sum(num); Q = Math.maximum(Q, 1e-12); # Compute gradient PQ = P - Q; for i in range(n): dY[i,:] = Math.sum(Math.tile(PQ[:,i] * num[:,i], (no_dims, 1)).T * (Y[i,:] - Y), 0); # Perform the update if iter < 20: momentum = initial_momentum else: momentum = final_momentum gains = (gains + 0.2) * ((dY > 0) != (iY > 0)) + (gains * 0.8) * ((dY > 0) == (iY > 0)); gains[gains < min_gain] = min_gain; iY = momentum * iY - eta * (gains * dY); Y = Y + iY; Y = Y - Math.tile(Math.mean(Y, 0), (n, 1)); # Compute current value of cost function if (iter + 1) % 10 == 0: C = Math.sum(P * Math.log(P / Q)); print "Iteration ", (iter + 1), ": error is ", C # Stop lying about P-values if iter == 100: P = P / 4; # Return solution return Y; if __name__ == "__main__": print "Run Y = tsne.tsne(X, no_dims, perplexity) to perform t-SNE on your dataset." print "Running example on 2,500 MNIST digits..." X = Math.loadtxt("train_X4000.txt");#X = X[:100] labels = Math.loadtxt("train_y4000.txt");#labels = labels[:100] Y = tsne(X, 2, 38, 20.0); fil = open('Y.txt','w') for i in Y: fil.write(str(i[0])+' '+str(i[1])+'\n') fil.close() colors=['b', 'c', 'y', 'm', 'r'] idx_1 = [i1 for i1 in range(len(labels)) if labels[i1]==1] flg1=Plot.scatter(Y[idx_1,0], Y[idx_1,1], 20,color=colors[0],label='1'); idx_2= [i2 for i2 in range(len(labels)) if labels[i2]==2] flg2=Plot.scatter(Y[idx_2,0], Y[idx_2,1], 20,color=colors[1], label='2'); idx_3= [i3 for i3 in range(len(labels)) if labels[i3]==3] flg3=Plot.scatter(Y[idx_3,0], Y[idx_3,1], 20, color=colors[2],label='3'); idx_4= [i4 for i4 in range(len(labels)) if labels[i4]==4] flg4=Plot.scatter(Y[idx_4,0], Y[idx_4,1], 20,color=colors[3], label='4'); idx_5= [i5 for i5 in range(len(labels)) if labels[i5]==5] flg5=Plot.scatter(Y[idx_5,0], Y[idx_5,1], 20, color=colors[4],label='5');# flg=Plot.scatter(Y[:,0], Y[:,1], 20,labels); Plot.legend() Plot.savefig('figure4000.pdf') Plot.show() |





【推荐】国内首个AI IDE,深度理解中文开发场景,立即下载体验Trae
【推荐】编程新体验,更懂你的AI,立即体验豆包MarsCode编程助手
【推荐】抖音旗下AI助手豆包,你的智能百科全书,全免费不限次数
【推荐】轻量又高性能的 SSH 工具 IShell:AI 加持,快人一步
· 开发者必知的日志记录最佳实践
· SQL Server 2025 AI相关能力初探
· Linux系列:如何用 C#调用 C方法造成内存泄露
· AI与.NET技术实操系列(二):开始使用ML.NET
· 记一次.NET内存居高不下排查解决与启示
· 阿里最新开源QwQ-32B,效果媲美deepseek-r1满血版,部署成本又又又降低了!
· 开源Multi-agent AI智能体框架aevatar.ai,欢迎大家贡献代码
· Manus重磅发布:全球首款通用AI代理技术深度解析与实战指南
· 被坑几百块钱后,我竟然真的恢复了删除的微信聊天记录!
· 没有Manus邀请码?试试免邀请码的MGX或者开源的OpenManus吧