>>> ps -ef | grep filename 查看关于filename文件的所有进程状态
>>> find dirpath -name filename 在dirpath目录(绝对路径)下查找关于filename的所有文件
>>> fg %jobsnumber 将后台命令调用到前台运行
>>> Ctrl + z 挂起当前运行任务 -jobs filename number 查找运行文件filename的PID号 bg %(jobsnumber)将挂起的文件放在后台运行
>>> Ctrl + c 结束当前运行的任务
>>> cat score.sc | sort -nk2 > score_sorted.txt 对得分文件score.sc的第二列按照数值(n)的大小进行排序,并将排序好的信息存储在score_sorted.txt文件中。
>>> 对模板PDB文件进行处理: python rosetta_tools/protein_tools/scripts/clean_pdb.py 2anv.pdb A
>>> 基于 rosettaCM_1_3.py脚本 同源建模

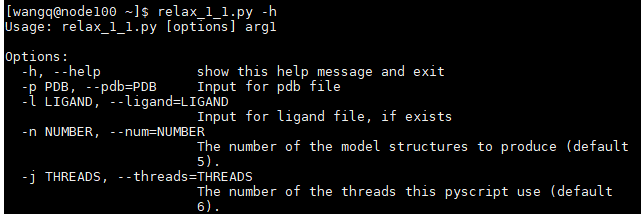
>>> 基于relax_1_1.py 脚本 模型优化
params文件的制备(对于对接小分子的.sdf mol 和mol2)
python rosetta_source/src/python/apps/public/molfile_to_params.py -n MR3 MR3.mol 得到两个文件 MR3.params and MR3_0001.pdb
dock.xml 文件的设置
<ROSETTASCRIPTS>
<SCOREFXNS>
<ligand_soft_rep weights="ligand_soft_rep">
</ligand_soft_rep>
<hard_rep weights="ligand">
</hard_rep>
</SCOREFXNS>
<LIGAND_AREAS>
<docking_sidechain_X chain="X" cutoff="6.0" add_nbr_radius="true" all_atom_mode="true" minimize_ligand="10"/>
<final_sidechain_X chain="X" cutoff="6.0" add_nbr_radius="true" all_atom_mode="true"/>
<final_backbone_X chain="X" cutoff="7.0" add_nbr_radius="false" all_atom_mode="true" Calpha_restraints="0.3"/>
<docking_sidechain_F chain="F" cutoff="6.0" add_nbr_radius="true" all_atom_mode="true" minimize_ligand="10"/>
<final_sidechain_F chain="F" cutoff="6.0" add_nbr_radius="true" all_atom_mode="true"/>
<final_backbone_F chain="F" cutoff="7.0" add_nbr_radius="false" all_atom_mode="true" Calpha_restraints="0.3"/>
</LIGAND_AREAS>
<INTERFACE_BUILDERS>
<side_chain_for_docking ligand_areas="docking_sidechain_X,docking_sidechain_F"/>
<side_chain_for_final ligand_areas="final_sidechain_X,final_sidechain_F"/>
<backbone ligand_areas="final_backbone_X,final_backbone_F" extension_window="3"/>
</INTERFACE_BUILDERS>
<MOVEMAP_BUILDERS>
<docking sc_interface="side_chain_for_docking" minimize_water="true"/>
<final sc_interface="side_chain_for_final" bb_interface="backbone" minimize_water="true"/>
</MOVEMAP_BUILDERS>
<FILTERS>
<AtomicDistance name="S-HEM" residue1="435A" atomtype1="S" residue2="1F" atomtype2="Fe3p" distance="2.60" confidence="0.9"/>
</FILTERS>
<SCORINGGRIDS ligand_chain="F" width="15">
<classic grid_type="ClassicGrid" weight="1.0"/>
</SCORINGGRIDS>
<MOVERS>
single movers_X
<Transform name="transform_F" chain="F" box_size="7.0" move_distance="0.2" angle="20" cycles="500" repeats="1" temperature="5"/>
<CompoundTranslate name="compound_translate" randomize_order="false" allow_overlap="false">
<Translate chain="X" distribution="uniform" angstroms="3.0" cycles="50"/>
<Translate chain="F" distribution="uniform" angstroms="3.0" cycles="50"/>
</CompoundTranslate>
<Rotate name="rotate_X" chain="X" distribution="uniform" degrees="360" cycles="700"/>
<Rotate name="rotate_F" chain="F" distribution="uniform" degrees="360" cycles="700"/>
<SlideTogether name="slide_together" chains="X,F"/>
<HighResDocker name="high_res_docker" cycles="6" repack_every_Nth="3" scorefxn="ligand_soft_rep" movemap_builder="docking"/>
<FinalMinimizer name="final" scorefxn="hard_rep" movemap_builder="final"/>
<InterfaceScoreCalculator name="add_scores" chains="X,F" scorefxn="hard_rep"/>
compound movers
<ParsedProtocol name="low_res_dock">
<Add mover_name="transform_F"/>
<Add mover_name="compound_translate"/>
<Add mover_name="rotate_X"/>
<Add mover_name="rotate_F"/>
<Add mover_name="slide_together"/>
</ParsedProtocol>
<ParsedProtocol name="high_res_dock">
<Add mover_name="high_res_docker"/>
<Add mover_name="final"/>
</ParsedProtocol>
</MOVERS>
<PROTOCOLS>
<Add mover_name="low_res_dock"/>
<Add mover_name="high_res_dock"/>
<Add filter="S-HEM" />
<Add mover_name="add_scores"/>
</PROTOCOLS>
</ROSETTASCRIPTS>
dock.options设置
-in:file:s inputs_AGI/EbF6H_HEM_AGI.pdb -in:file:extra_res_fa inputs_AGI/HEM.params inputs_AGI/AGI.params -packing -ex1 -ex2aro -ex2 -no_optH false -flip_HNQ true -ignore_ligand_chi true -parser -protocol inputs_AGI/ligand_dock.xml -out -path:all outputs_AGI1 -nstruct 10000 -overwrite
>>> 分子对接 ./rosetta/main/source/bin/rosetta_scripts.linuxgccrelease @dock.options -database /rosetta/main/database -nstruct 1000 构建1000个对接模型 由total_score和其他参数选取最优的对接模型
——————————————————Small-molecule ligand docking into comparative models with Rosett 一篇关于rosettaCM流程详细操作的文章


